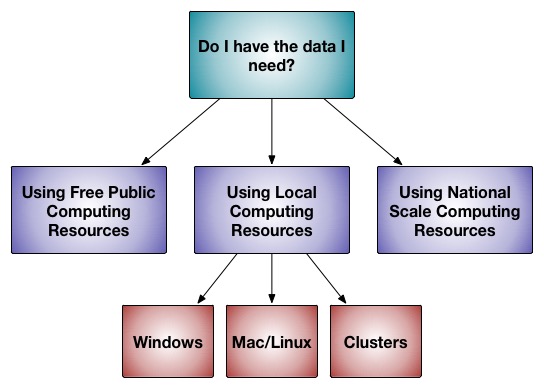
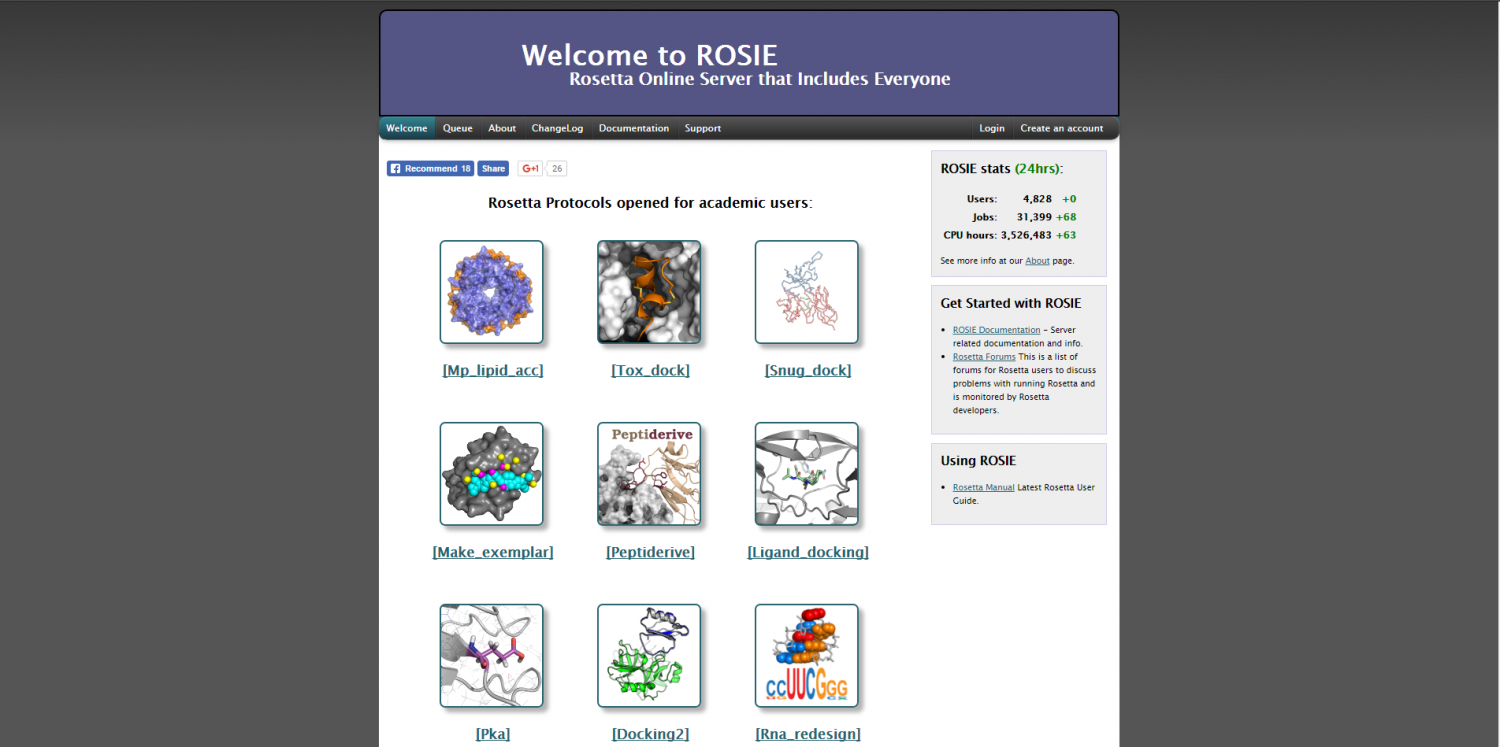
GalaxyDomDock: An Ab Initio Domain–domain Docking Web Server for Multi-domain Protein Structure Prediction - ScienceDirect
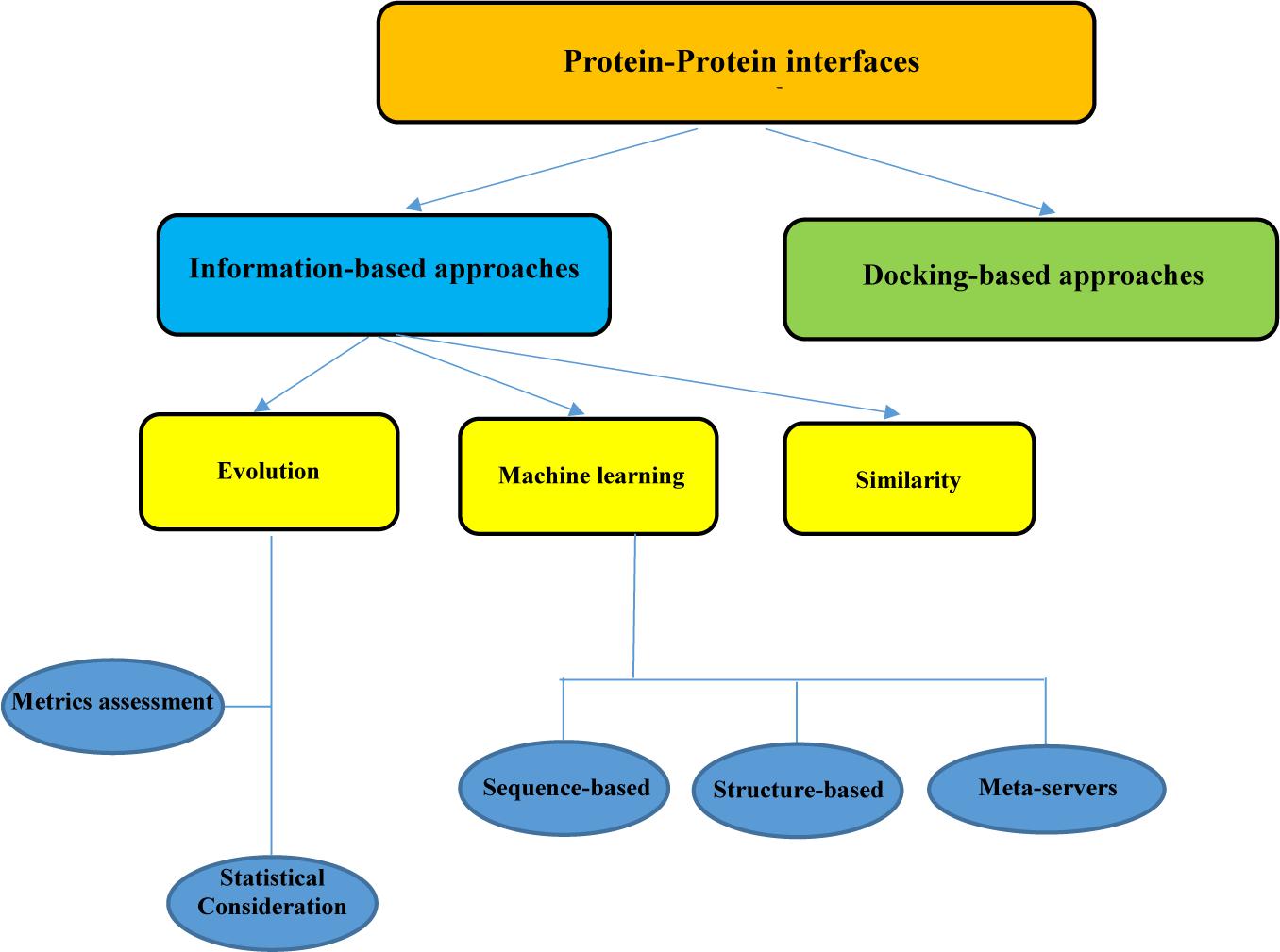
Frontiers | In silico Approaches for the Design and Optimization of Interfering Peptides Against Protein–Protein Interactions
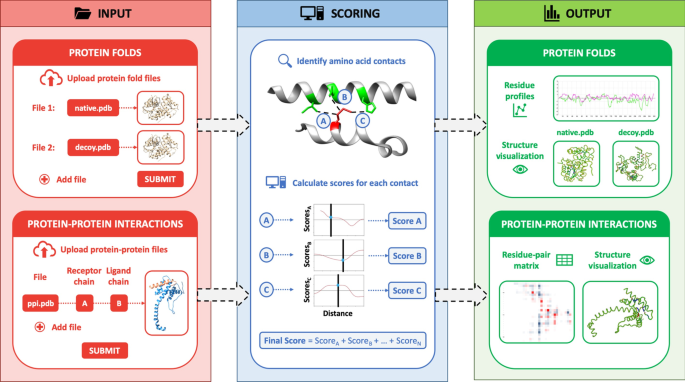
SPServer: split-statistical potentials for the analysis of protein structures and protein–protein interactions | BMC Bioinformatics | Full Text

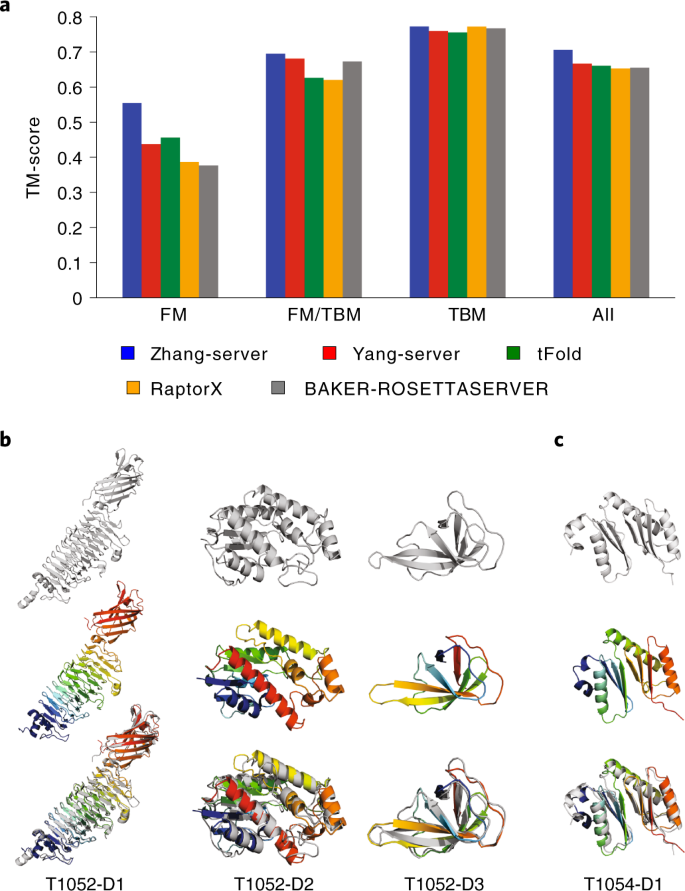
Accurate prediction of protein structures and interactions using a three-track neural network | Science
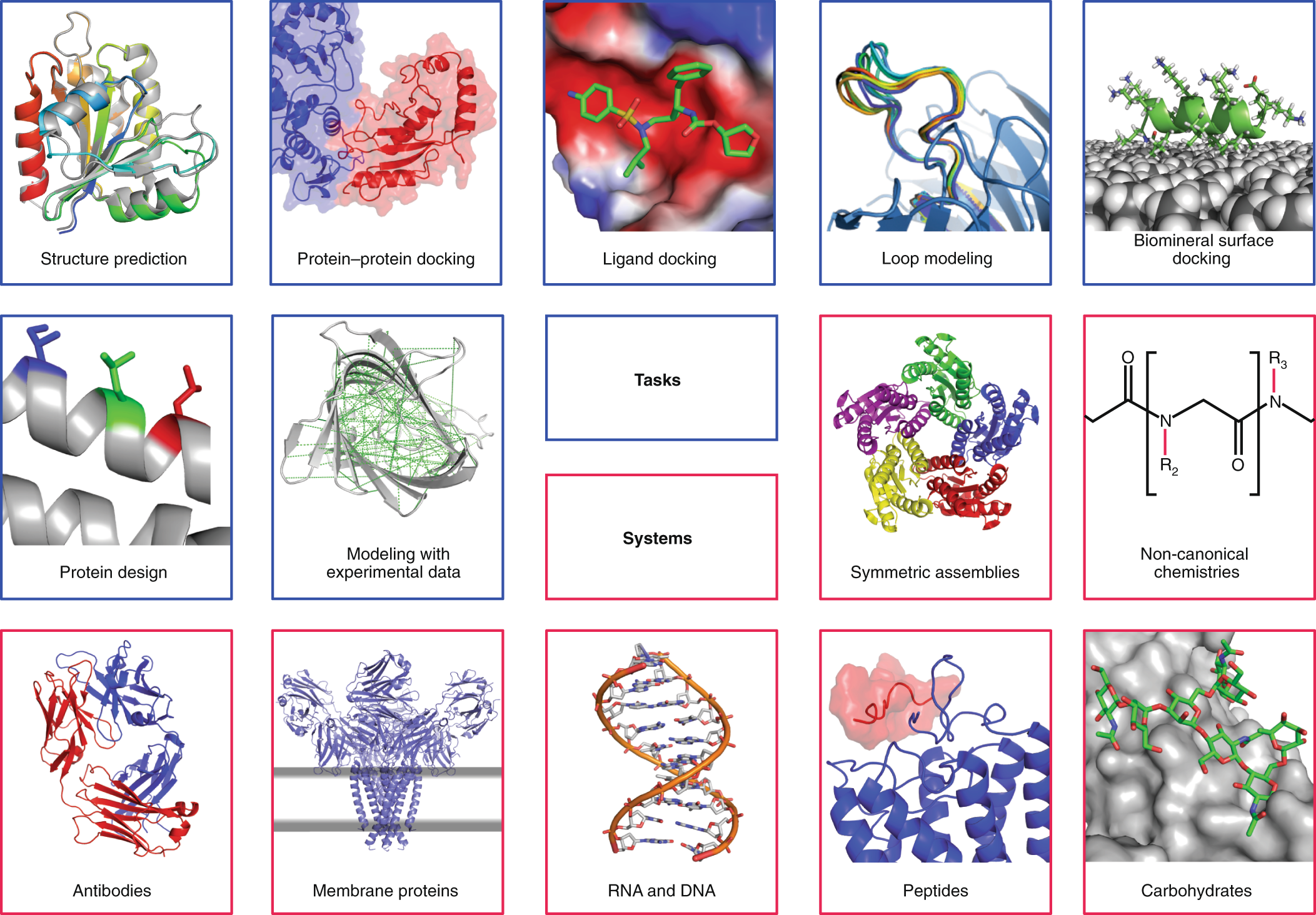
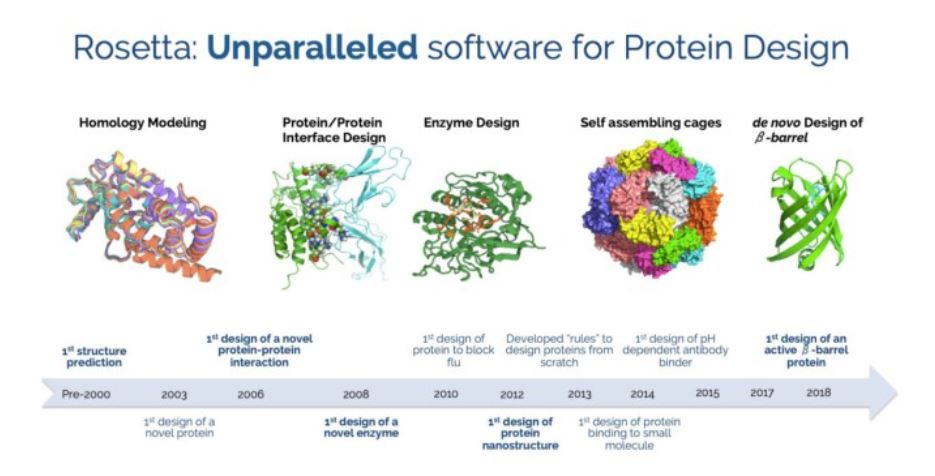
1 Macromolecular modeling and design in Rosetta: new methods and frameworks Koehler Leman, Julia [1, 2]*, Weitzner, Brian D [3,

Accurate prediction of protein structures and interactions using a three-track neural network | Science



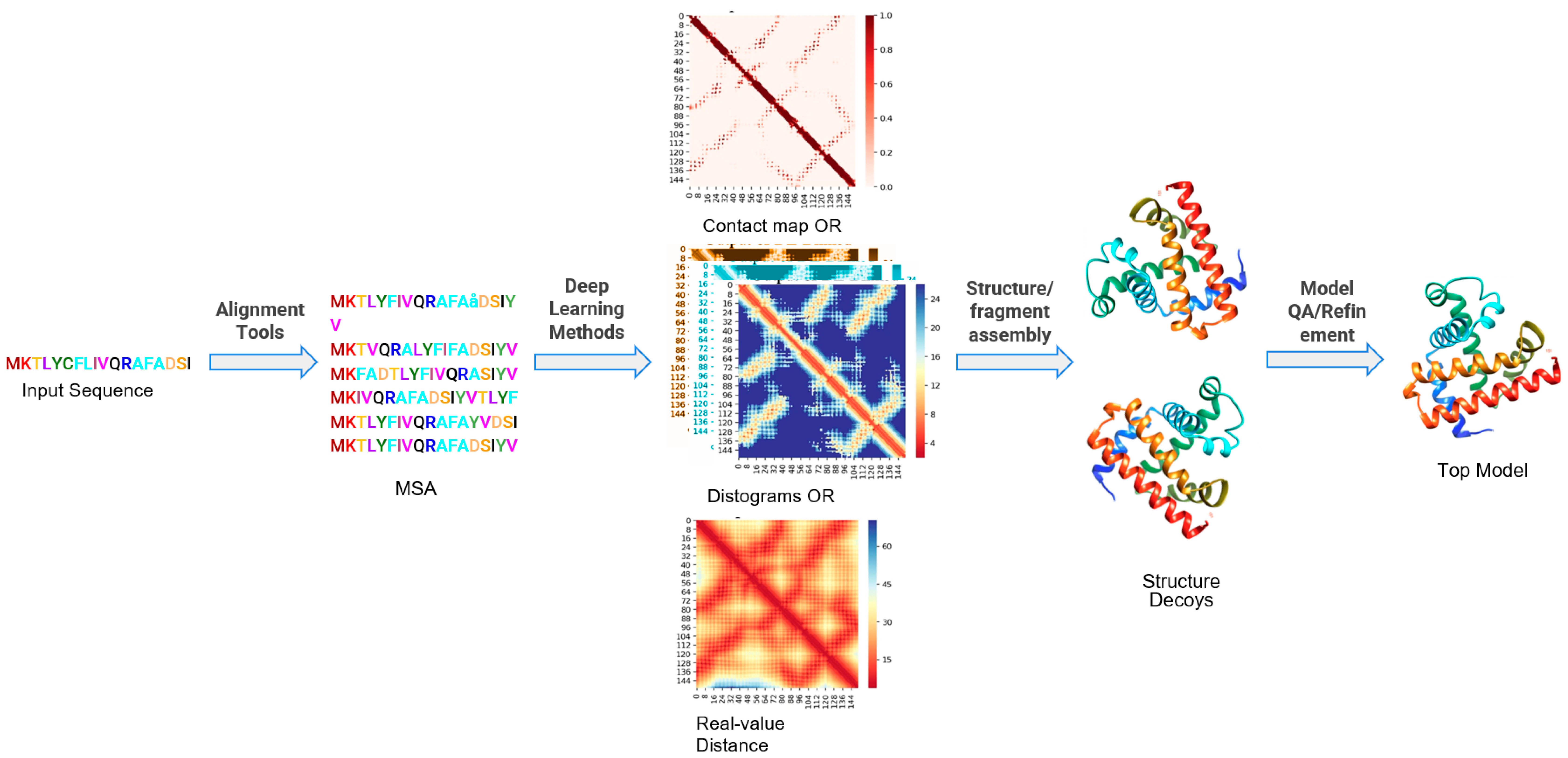
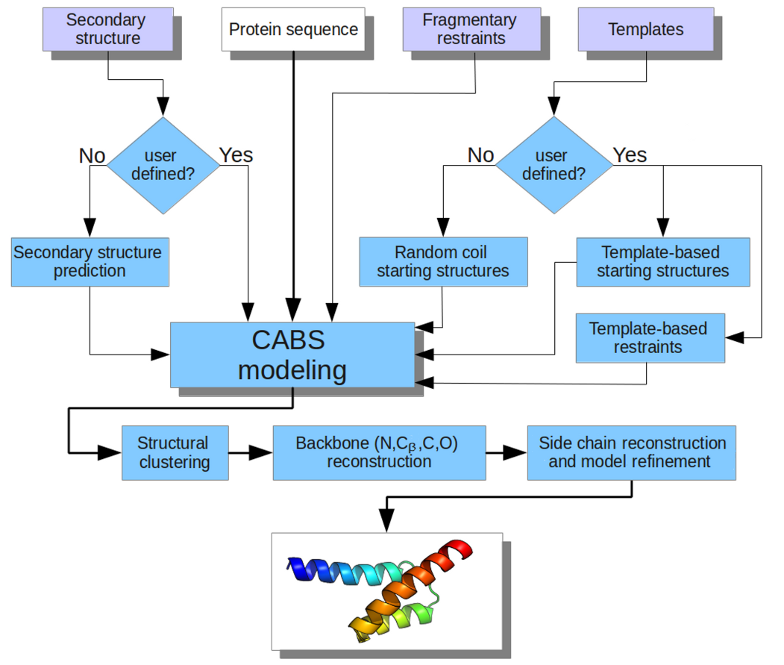
![General workflow of Rosetta for protein structure prediction [53] | Download Scientific Diagram General workflow of Rosetta for protein structure prediction [53] | Download Scientific Diagram](https://www.researchgate.net/profile/Theam-Lim/publication/281540998/figure/fig2/AS:281366399864835@1444094385469/General-workflow-of-Rosetta-for-protein-structure-prediction-53_Q640.jpg)
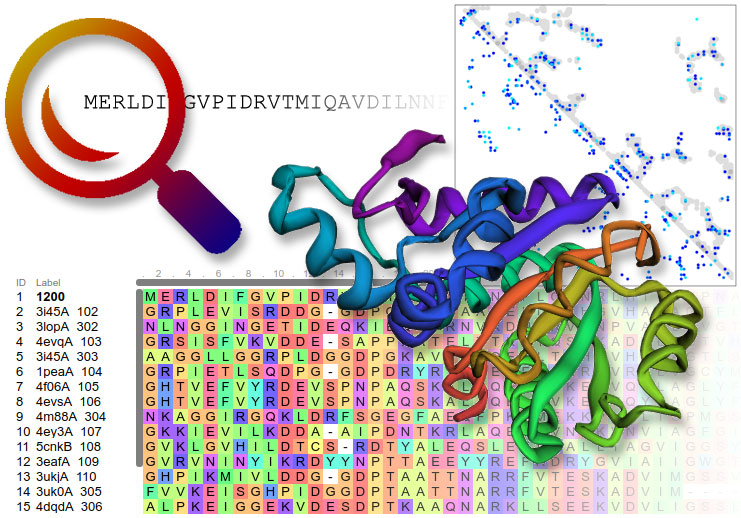




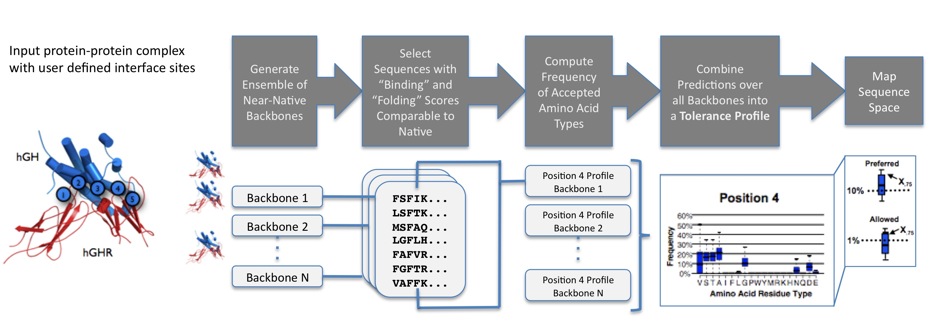
![PDF] Protein structure prediction and analysis using the Robetta server | Semantic Scholar PDF] Protein structure prediction and analysis using the Robetta server | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/0fa67c023e732439f6e0dc5f012e7875ea1cad0c/2-Figure1-1.png)