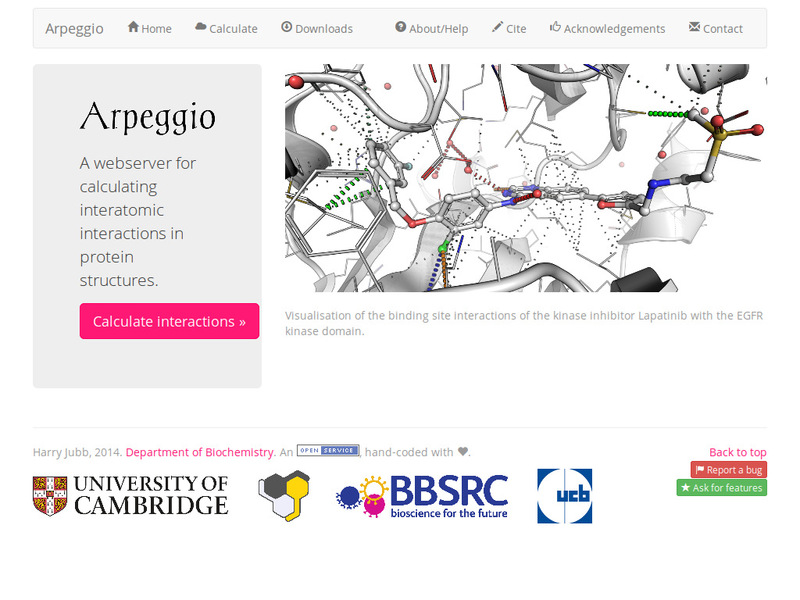
Arpeggio: A Web Server for Calculating and Visualising Interatomic Interactions in Protein Structures | Semantic Scholar

BINANA 2: Characterizing Receptor/Ligand Interactions in Python and JavaScript | Journal of Chemical Information and Modeling
fingeRNAt—A novel tool for high-throughput analysis of nucleic acid-ligand interactions | PLOS Computational Biology
Arpeggio: A web server for calculating and visualising interatomic interactions in protein structures
Arpeggio: A web server for calculating and visualising interatomic interactions in protein structures

Arpeggio: A Web Server for Calculating and Visualising Interatomic Interactions in Protein Structures - ScienceDirect

IJMS | Free Full-Text | PremPRI: Predicting the Effects of Missense Mutations on Protein–RNA Interactions
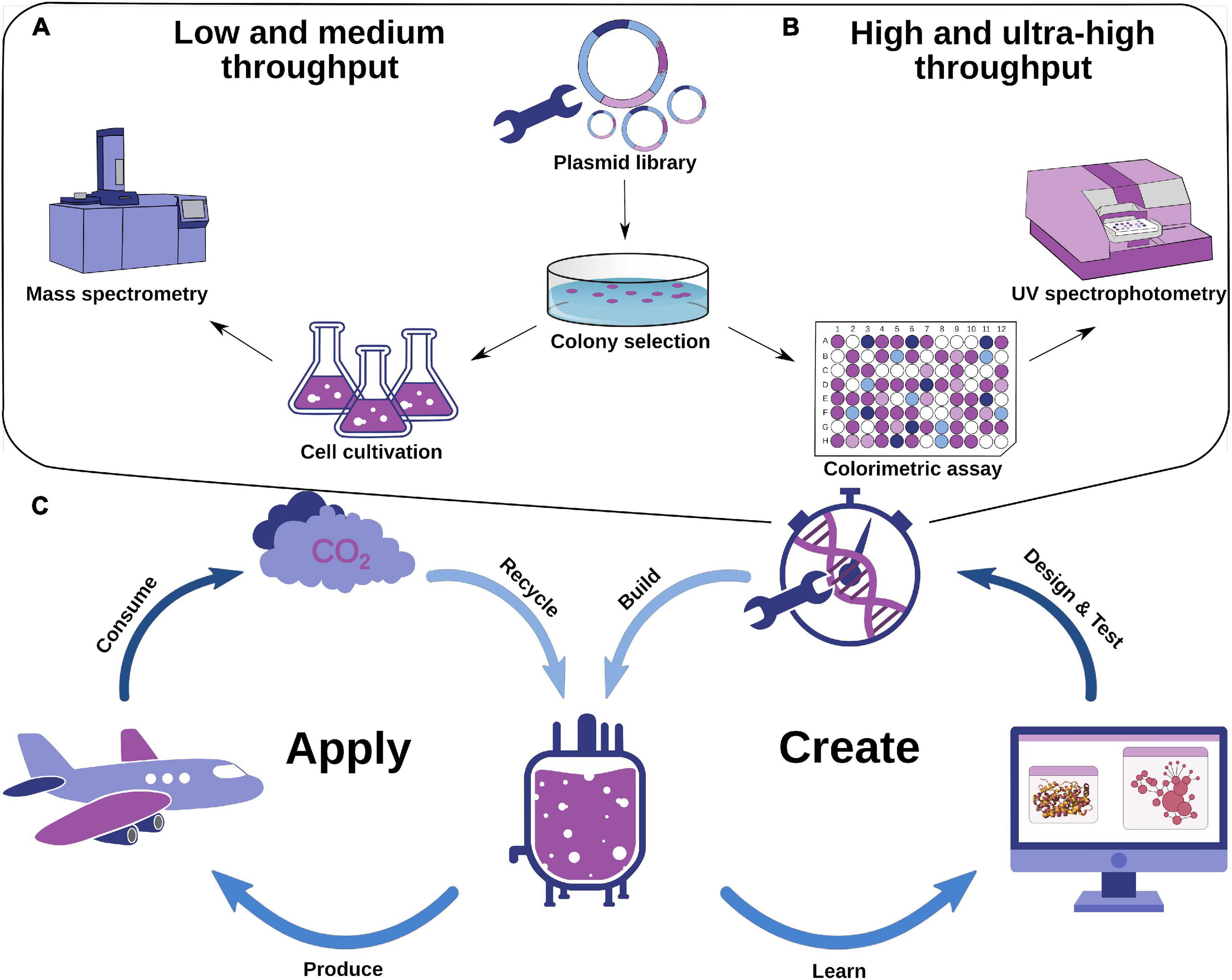
Frontiers | Computational Enzyme Engineering Pipelines for Optimized Production of Renewable Chemicals

A Comprehensive Computational Platform to Guide Drug Development Using Graph-Based Signature Methods | SpringerLink

PDF) Arpeggio: A Web Server for Calculating and Visualising Interatomic Interactions in Protein Structures
Arpeggio: A Web Server for Calculating and Visualising Interatomic Interactions in Protein Structures

Molecules | Free Full-Text | Infection Dynamics of ATG8 in Leishmania: Balancing Autophagy for Therapeutics

PDF) Arpeggio: A Web Server for Calculating and Visualising Interatomic Interactions in Protein Structures

BINANA 2: Characterizing Receptor/Ligand Interactions in Python and JavaScript | Journal of Chemical Information and Modeling

.png.1b55a634569ab2467e3341ccbad99a48.png)